GeneSurfer enables transcriptome-wide exploration and annotation of gene co-expression modules in 3D spatial transcriptomics data
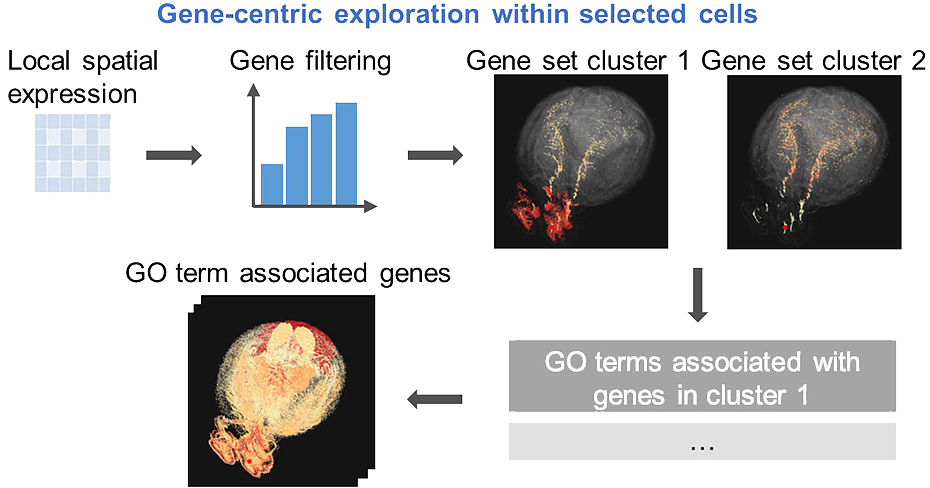
Gene co-expression provides crucial insights into biological functions, however, there is a lack of exploratory analysis tools for localized gene co-expression in large-scale datasets. We present GeneSurfer, an interactive interface designed to explore localized transcriptome-wide gene co-expression patterns in the 3D spatial domain. Key features of GeneSurfer include transcriptome-wide gene filtering and gene clustering based on spatial local co-expression within transcriptomically similar cells, multi-slice 3D rendering of average expression of gene clusters, and on-the-fly Gene Ontology term annotation of co-expressed gene sets. Additionally, GeneSurfer offers multiple linked views for investigating individual genes or gene co-expression in the spatial domain at each exploration stage. Demonstrating its utility with both spatially resolved transcriptomics and single-cell RNA sequencing data from the Allen Brain Cell Atlas, GeneSurfer effectively identifies and annotates localized transcriptome-wide co-expression, providing biological insights and facilitating hypothesis generation and validation.
Resources
Citation
BibTeX
@article{ bib:2025_genesurfer,
author = {Alexander Vieth and Anna Vilanova and Boudewijn Lelieveldt and Elmar Eisemann and Thomas H{\"o}llt},
title = { GeneSurfer enables transcriptome-wide exploration and annotation of gene co-expression modules in 3D spatial transcriptomics data },
journal = { iScience },
pages = { 112713 },
year = { 2025 },
doi = { 10.1016/j.isci.2025.112713 },
}