Cytosplore EvoViewer: Visual Analytics of Conserved Evolutionary Patterns in multi-species single-cell sequencing data
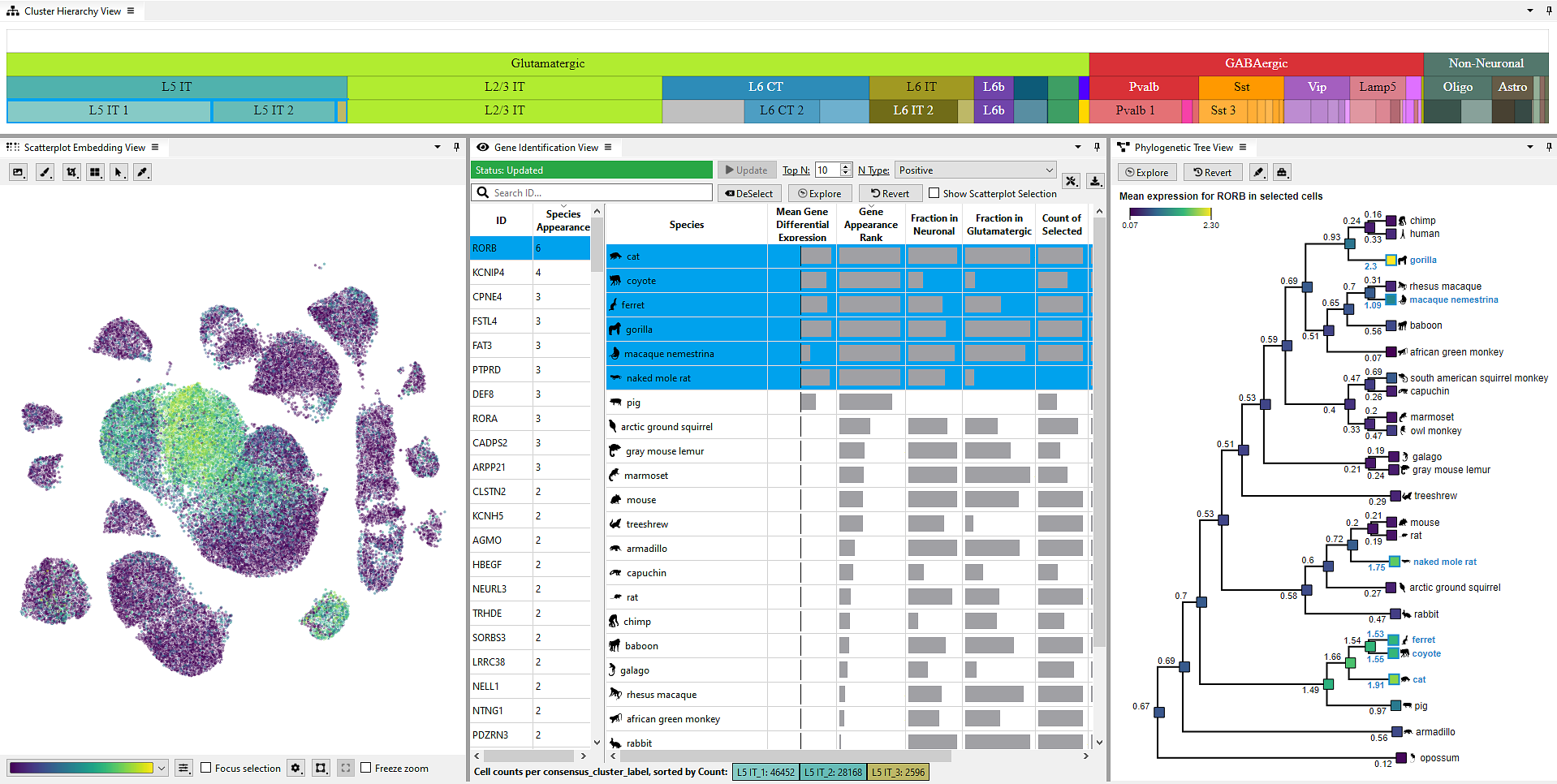
Single-cell transcriptomics has enhanced our understanding of the brain’s cellular composition. Biologists now analyze complex datasets to explore how marker genes influence biological processes, genetic variations, and phenotypic traits. A challenge is comparing these datasets across species to detect subtle differences or similarities to evolutionary development. Here, we present Cytosplore EvoViewer to facilitate examining relationships between transcriptomic datasets across species, simplifying the analysis of marker gene regulation and its impact on biological functions and integrating these findings with prior evolutionary knowledge or species-specific traits. We conducted a design study, including domain analysis, implementation of the results into Cytosplore EvoViewer, and an expert evaluation. Cytosplore EvoViewer offers valuable insights into genetic variations and evolutionary dynamics, helping to understand the diversity and the unity within diversity across species and their evolutionary development.
Resources
Citation
BibTeX
@inproceedings{ bib:2025_evoviewer,
author = {vibasueth and Morgan Wirthlin and Jeroen Eggermont and Thomas Kroes and Boudewijn Lelieveldt and Ed S. Lein and Trygve E. Bakken and Thomas H{\"o}llt},
title = { Cytosplore EvoViewer: Visual Analytics of Conserved Evolutionary Patterns in multi-species single-cell sequencing data },
booktitle = { Proceedings of the 18th IEEE Pacific Visualization Symposium },
pages = { 149 -- 159 },
year = { 2025 },
doi = { 10.1109/PacificVis64226.2025.00021 },
}